Rat PS1 (Presenilin 1) CLIA Kit (RTES00629)
- SKU:
- RTES00629
- Product Type:
- ELISA Kit
- ELISA Type:
- CLIA Kit
- Size:
- 96 Assays
- Sensitivity:
- 18.75pg/mL
- Range:
- 31.25-2000pg/mL
- ELISA Type:
- Sandwich
- Reactivity:
- Rat
- Sample Type:
- Serum, plasma and other biological fluids
- Research Area:
- Cell Death
Description
| Assay type: | Sandwich |
| Format: | 96T |
| Assay time: | 4.5h |
| Reactivity: | Rat |
| Detection method: | Chemiluminescence |
| Detection range: | 31.25-2000 pg/mL |
| Sensitivity: | 18.75 pg/mL |
| Sample volume: | 100µL |
| Sample type: | Serum, plasma and other biological fluids |
| Repeatability: | CV < 15% |
| Specificity: | This kit recognizes Rat PS1 in samples. No significant cross-reactivity or interference between Rat PS1 and analogues was observed. |
This kit uses Sandwich-CLIA as the method. The micro CLIA plate provided in this kit has been pre-coated with an antibody specific to Rat PS1. Standards or samples are added to the appropriate micro CLIA plate wells and combined with the specific antibody. Then a biotinylated detection antibody specific for Rat PS1 and Avidin-Horseradish Peroxidase (HRP) conjugate are added to each micro plate well successively and incubated. Free components are washed away. The substrate solution is added to each well. Only those wells that contain Rat PS1, biotinylated detection antibody and Avidin-HRP conjugate will appear fluorescence. The Relative light unit (RLU) value is measured spectrophotometrically by the Chemiluminescence immunoassay analyzer. The RLU value is positively associated with the concentration of Rat PS1. The concentration of Rat PS1 in the samples can be calculated by comparing the RLU of the samples to the standard curve.
| UniProt Protein Function: | PSEN1: probable catalytic subunit of the gamma-secretase complex, an endoprotease complex that catalyzes the intramembrane cleavage of integral membrane proteins such as Notch receptors and APP (beta-amyloid precursor protein). Requires the other members of the gamma-secretase complex to have a protease activity. May play a role in intracellular signaling and gene expression or in linking chromatin to the nuclear membrane. Regulates epithelial- cadherin function. Five alternative splice isoforms have been identified. |
| UniProt Protein Details: | Protein type:Membrane protein, integral; EC 3. 4. 23. -; Cell surface; Mitochondrial; Membrane protein, multi-pass; Protease Chromosomal Location of Human Ortholog: 14q24. 3 Cellular Component: Golgi apparatus; nuclear outer membrane; centrosome; cell surface; smooth endoplasmic reticulum; mitochondrion; integral to plasma membrane; lysosomal membrane; integral to membrane; lipid raft; kinetochore; ciliary rootlet; cell soma; membrane; perinuclear region of cytoplasm; mitochondrial inner membrane; apical plasma membrane; cytoplasmic vesicle; dendritic shaft; neuromuscular junction; endoplasmic reticulum membrane; nuclear membrane; rough endoplasmic reticulum; endoplasmic reticulum; cell cortex; Z disc; Golgi membrane; growth cone; axon Molecular Function:protein binding; cadherin binding; calcium channel activity; endopeptidase activity; beta-catenin binding; aspartic-type endopeptidase activity; PDZ domain binding Biological Process: positive regulation of catalytic activity; extracellular matrix organization and biogenesis; activation of MAPKK activity; positive regulation of apoptosis; beta-amyloid formation; regulation of synaptic plasticity; positive regulation of coagulation; Wnt receptor signaling pathway through beta-catenin; choline transport; T cell receptor signaling pathway; mitochondrial transport; post-embryonic development; T cell activation during immune response; extracellular matrix disassembly; positive regulation of MAP kinase activity; epithelial cell proliferation; cell-cell adhesion; negative regulation of axonogenesis; negative regulation of neuron apoptosis; embryonic limb morphogenesis; autophagic vacuole formation; somitogenesis; regulation of phosphorylation; skin morphogenesis; Notch receptor processing; negative regulation of ubiquitin-protein ligase activity; neuron development; Cajal-Retzius cell differentiation; hemopoietic progenitor cell differentiation; response to oxidative stress; negative regulation of apoptosis; regulation of synaptic transmission, glutamatergic; protein maturation; neuron migration; protein amino acid glycosylation; cell fate specification; myeloid dendritic cell differentiation; negative regulation of transcription from RNA polymerase II promoter; protein transport; amyloid precursor protein catabolic process; beta-amyloid metabolic process; regulation of resting membrane potential; brain morphogenesis; heart looping; positive regulation of receptor recycling; dorsoventral neural tube patterning; blood vessel development; Notch signaling pathway; L-glutamate transport; thymus development; membrane protein ectodomain proteolysis; memory; negative regulation of epidermal growth factor receptor activity; smooth endoplasmic reticulum calcium ion homeostasis; neuron apoptosis; endoplasmic reticulum calcium ion homeostasis; cerebral cortex cell migration; skeletal morphogenesis; positive regulation of proteasomal ubiquitin-dependent protein catabolic process; regulation of protein binding; protein processing; synaptic vesicle targeting; response to DNA damage stimulus Disease: Alzheimer Disease 3; Pick Disease Of Brain; Frontotemporal Dementia; Acne Inversa, Familial, 3; Cardiomyopathy, Dilated, 1u |
| NCBI Summary: | Alzheimer's disease (AD) patients with an inherited form of the disease carry mutations in the presenilin proteins (PSEN1; PSEN2) or in the amyloid precursor protein (APP). These disease-linked mutations result in increased production of the longer form of amyloid-beta (main component of amyloid deposits found in AD brains). Presenilins are postulated to regulate APP processing through their effects on gamma-secretase, an enzyme that cleaves APP. Also, it is thought that the presenilins are involved in the cleavage of the Notch receptor, such that they either directly regulate gamma-secretase activity or themselves are protease enzymes. Several alternatively spliced transcript variants encoding different isoforms have been identified for this gene, the full-length nature of only some have been determined. [provided by RefSeq, Aug 2008] |
| UniProt Code: | P97887 |
| NCBI GenInfo Identifier: | 4506163 |
| NCBI Gene ID: | 5663 |
| NCBI Accession: | NP_000012. 1 |
| UniProt Secondary Accession: | P97887,P97887, P49769, Q8HXW5, |
| UniProt Related Accession: | P49768 |
| Molecular Weight: | 52668 |
| NCBI Full Name: | presenilin-1 isoform I-467 |
| NCBI Synonym Full Names: | presenilin 1 |
| NCBI Official Symbol: | PSEN1 |
| NCBI Official Synonym Symbols: | AD3; FAD; PS1; PS-1; S182; ACNINV3 |
| NCBI Protein Information: | presenilin-1 |
| UniProt Protein Name: | Presenilin-1 |
| UniProt Synonym Protein Names: | Protein S182 |
| UniProt Gene Name: | PSEN1 |
| UniProt Entry Name: | PSN1_HUMAN |
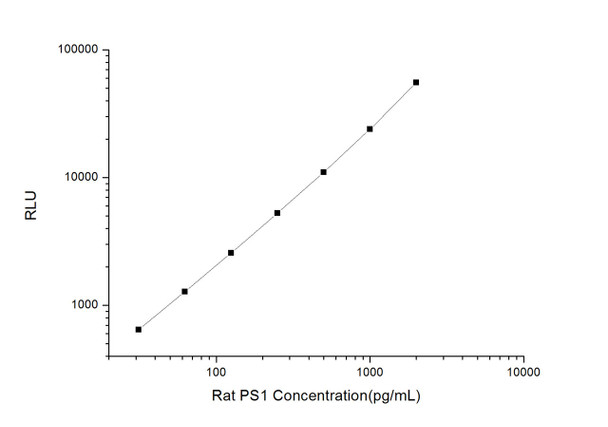
As the RLU values of the standard curve may vary according to the conditions of the actual assay performance (e. g. operator, pipetting technique, washing technique or temperature effects), the operator should establish a standard curve for each test. Typical standard curve and data is provided below for reference only.
| Concentration (pg/mL) | RLU | Average | Corrected |
| 2000 | 50618 60824 | 55721 | 55693 |
| 1000 | 22698 25250 | 23974 | 23946 |
| 500 | 11111 10953 | 11032 | 11004 |
| 250 | 4890 5698 | 5294 | 5266 |
| 125 | 2655 2561 | 2608 | 2580 |
| 62.5 | 1328 1294 | 1311 | 1283 |
| 31.25 | 656 692 | 674 | 646 |
| 0 | 27 29 | 28 | -- |
Precision
Intra-assay Precision (Precision within an assay): 3 samples with low, mid range and high level Rat PS1 were tested 20 times on one plate, respectively.
Inter-assay Precision (Precision between assays): 3 samples with low, mid range and high level Rat PS1 were tested on 3 different plates, 20 replicates in each plate.
| Intra-assay Precision | Inter-assay Precision | |||||
| Sample | 1 | 2 | 3 | 1 | 2 | 3 |
| n | 20 | 20 | 20 | 20 | 20 | 20 |
| Mean (pg/mL) | 102.96 | 256.54 | 990.62 | 105.84 | 246.68 | 922.03 |
| Standard deviation | 9.20 | 23.24 | 61.22 | 12.79 | 25.46 | 73.58 |
| C V (%) | 8.94 | 9.06 | 6.18 | 12.08 | 10.32 | 7.98 |
Recovery
The recovery of Rat PS1 spiked at three different levels in samples throughout the range of the assay was evaluated in various matrices.
| Sample Type | Range (%) | Average Recovery (%) |
| Serum (n=5) | 87-101 | 95 |
| EDTA plasma (n=5) | 95-112 | 102 |
| Cell culture media (n=5) | 96-111 | 102 |
Linearity
Samples were spiked with high concentrations of Rat PS1 and diluted with Reference Standard & Sample Diluent to produce samples with values within the range of the assay.
| Serum (n=5) | EDTA plasma (n=5) | Cell culture media (n=5) | ||
| 1:2 | Range (%) | 97-114 | 97-111 | 91-106 |
| Average (%) | 105 | 104 | 99 | |
| 1:4 | Range (%) | 87-101 | 87-103 | 92-106 |
| Average (%) | 94 | 94 | 99 | |
| 1:8 | Range (%) | 95-111 | 97-111 | 95-108 |
| Average (%) | 103 | 103 | 102 | |
| 1:16 | Range (%) | 97-111 | 100-115 | 94-109 |
| Average (%) | 104 | 106 | 100 |
An unopened kit can be stored at 4°C for 1 month. If the kit is not used within 1 month, store the items separately according to the following conditions once the kit is received.
| Item | Specifications | Storage |
| Micro CLIA Plate(Dismountable) | 8 wells ×12 strips | -20°C, 6 months |
| Reference Standard | 2 vials | |
| Concentrated Biotinylated Detection Ab (100×) | 1 vial, 120 µL | |
| Concentrated HRP Conjugate (100×) | 1 vial, 120 µL | -20°C(shading light), 6 months |
| Reference Standard & Sample Diluent | 1 vial, 20 mL | 4°C, 6 months |
| Biotinylated Detection Ab Diluent | 1 vial, 14 mL | |
| HRP Conjugate Diluent | 1 vial, 14 mL | |
| Concentrated Wash Buffer (25×) | 1 vial, 30 mL | |
| Substrate Reagent A | 1 vial, 5 mL | 4°C (shading light) |
| Substrate Reagent B | 1 vial, 5 mL | 4°C (shading light) |
| Plate Sealer | 5 pieces | |
| Product Description | 1 copy | |
| Certificate of Analysis | 1 copy |
- Set standard, test sample and control (zero) wells on the pre-coated plate and record theirpositions. It is recommended to measure each standard and sample in duplicate. Note: addall solutions to the bottom of the plate wells while avoiding contact with the well walls. Ensuresolutions do not foam when adding to the wells.
- Aliquot 100µl of standard solutions into the standard wells.
- Add 100µl of Sample / Standard dilution buffer into the control (zero) well.
- Add 100µl of properly diluted sample (serum, plasma, tissue homogenates and otherbiological fluids. ) into test sample wells.
- Cover the plate with the sealer provided in the kit and incubate for 90 min at 37°C.
- Aspirate the liquid from each well, do not wash. Immediately add 100µL of BiotinylatedDetection Ab working solution to each well. Cover the plate with a plate seal and gently mix. Incubate for 1 hour at 37°C.
- Aspirate or decant the solution from the plate and add 350µL of wash buffer to each welland incubate for 1-2 minutes at room temperature. Aspirate the solution from each well andclap the plate on absorbent filter paper to dry. Repeat this process 3 times. Note: a microplatewasher can be used in this step and other wash steps.
- Add 100µL of HRP Conjugate working solution to each well. Cover with a plate seal andincubate for 30 min at 37°C.
- Aspirate or decant the solution from each well. Repeat the wash process for five times asconducted in step 7.
- Add 100µL of Substrate mixture solution to each well. Cover with a new plate seal andincubate for no more than 5 min at 37°C. Protect the plate from light.
- Determine the RLU value of each well immediately.