Immunopeptidomics: Mapping the Hidden Landscape of the Cancer Immunome
Imagine a high-stakes game of hide-and-seek where the seeker is your immune system and the hider is a master of disguise: cancer. For decades, we believed the 'wanted' posters for tumor cells were limited to a few well-known mutations. However, a revolutionary field called immunopeptidomics is revealing that the surface of a cancer cell is actually covered in thousands of hidden messages, waiting to be decoded by the next generation of precision medicine.
Introduction
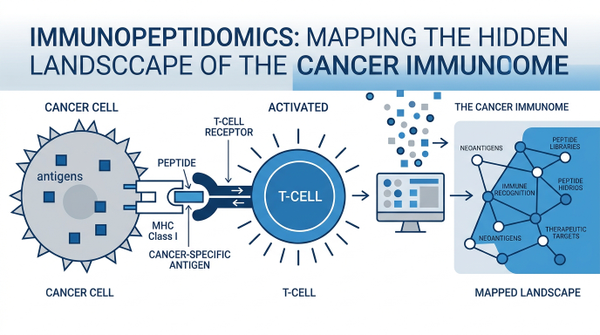
The human immune system relies on a sophisticated surveillance mechanism to distinguish 'self' from 'non-self.' At the heart of this process are Major Histocompatibility Complex (MHC) molecules, which act as cellular display cases, presenting small protein fragments called peptides to passing T cells. In a healthy cell, these peptides are routine; in a cancerous one, they should ideally signal distress. Yet, traditional genomics often misses the full picture because not every mutation results in a visible peptide, and not every visible peptide comes from a known gene. This is where immunopeptidomics steps in, utilizing ultra-sensitive mass spectrometry to directly sequence the peptides actually sitting on the cell surface. By mapping this 'hidden landscape,' researchers are moving beyond theoretical predictions to see the pan-cancer atlas reveals shared personalized neoantigen targets that were previously invisible. This shift from predicting what *might* be presented to seeing what *is* presented is fundamentally changing how we design vaccines and cell therapies, offering a direct look into the cancer immunome's most guarded secrets.
The Dark Matter of the Genome: Unmasking Cryptic Antigens
For years, the 'central dogma' of biology suggested that proteins only come from clearly defined genes. However, cancer cells are notorious for breaking the rules of biology. Recent breakthroughs have shown that a significant portion of the peptides presented on MHC molecules actually originate from 'non-coding' regions of the genome—areas previously dismissed as 'junk DNA.' These are known as cryptic antigens, and they represent a massive, untapped reservoir for immunotherapy. Interestingly, these antigens are often highly specific to tumor cells because the chaotic environment of a cancer cell allows for 'noisy' translation that healthy cells would never permit. Beyond this, researchers have found that cryptic antigens as targets for T-cell recognition are particularly prevalent in difficult-to-treat malignancies like pancreatic cancer. This discovery suggests that the 'hidden' part of the immunome might actually be the most vulnerable, providing a new set of targets for patients who currently have few options.
Why Cryptic Antigens are Game-Changers
The identification of these non-canonical targets shifts the focus from a few common mutations to a vast landscape of tumor-specific markers.
- Higher specificity: Many cryptic peptides are never expressed in normal tissues, reducing the risk of off-target effects.
- Shared targets: Some cryptic antigens are surprisingly common across different patients with the same cancer type.
- Overcoming resistance: Targeting multiple cryptic antigens makes it harder for tumors to 'hide' via single-gene mutations.
Decoding the Chaos: How Dysregulated Translation Feeds the Immunome
To understand why cancer cells present such a strange array of peptides, we have to look at the protein-making machinery itself. In a malignant state, the process of translation—turning RNA into protein—becomes fundamentally broken. This 'translation dysregulation' isn't just a byproduct of cancer; it is a source of its own destruction. When the cell's ribosomes start misreading the genetic code, they produce 'aberrant' peptides that the immune system can recognize as foreign. Recent studies in high-impact journals have confirmed that dysregulated translation generates novel targetable tumor antigens that are entirely unique to the tumor's metabolic state. This means that even if a tumor doesn't have a high 'mutational burden' in its DNA, it might still be highly visible to the immune system because of its messy internal chemistry. This finding matters because it expands the pool of patients who might benefit from immunotherapy, including those with 'cold' tumors that were previously thought to be invisible to T cells.
The Ribosome as a Target Discovery Tool
By studying how ribosomes stall or slip during translation, scientists can predict which 'errors' will end up on the cell surface as antigens.
From Data to Delivery: Building the Personalized Vaccine Pipeline
The ultimate goal of mapping the immunome is to create treatments tailored to the individual. However, the sheer volume of data generated by immunopeptidomics is overwhelming. To bridge the gap between the lab and the clinic, researchers are now integrating mass spectrometry with advanced computational models. This 'proteogenomic' approach allows for the rapid identification of the most immunogenic peptides from a patient's biopsy. For instance, a new proteogenomic pipeline optimizes personalized neoantigen discovery by filtering out the noise and focusing on the peptides most likely to trigger a strong T-cell response. This is particularly crucial for developing mRNA-based vaccines, where the choice of which 'message' to include determines the success of the therapy. Interestingly, immunopeptidomics identifies antigens for mRNA-based myeloma vaccines, demonstrating that this technology is already being used to design next-generation treatments for blood cancers. The speed and accuracy of these pipelines are what will finally make 'real-time' personalized oncology a reality.
The Multi-Omic Advantage
Combining different layers of biological data provides a 360-degree view of the tumor.
- Genomics: Identifies the potential mutations in the DNA.
- Transcriptomics: Confirms which genes are actually being turned into RNA.
- Immunopeptidomics: Proves which peptides are physically present on the MHC molecules.
Precision in Practice: Deep-Data Insights from the Front Lines
While the theory of immunopeptidomics is compelling, its real-world application requires overcoming significant technical hurdles, especially in solid tumors like glioblastoma. The brain is a notoriously difficult environment for both diagnosis and treatment. However, new techniques are allowing researchers to extract massive amounts of information from tiny tissue samples. Recent evidence shows that needle biopsies enable deep multi-omic glioblastoma analysis, providing a window into the immunopeptidome of the most aggressive brain cancers. Beyond this, the integration of full-length transcriptomics identifies diverse immunogenic neoantigens that were previously missed by shorter sequencing methods. This level of precision ensures that no potential target is left behind. By combining these deep-data approaches, we are finally moving toward a future where a single biopsy can provide the entire blueprint for a patient's personalized cure, mapping every hidden corner of the cancer immunome.
The Future of the Decoded Immunome
Immunopeptidomics has fundamentally shifted our perspective on cancer. We no longer see the tumor surface as a static barrier, but as a dynamic, information-rich landscape that tells the story of the cell's internal state. By unmasking cryptic antigens and leveraging the chaos of dysregulated translation, we are finding 'wanted' posters for cancer in places we never thought to look. As proteogenomic pipelines become faster and more integrated into clinical workflows, the dream of truly personalized immunotherapy is moving within reach. The hidden landscape of the cancer immunome is finally being mapped, and in those maps, we are finding the paths to a new era of precision medicine.
References
- Ely ZA, Kulstad ZJ, et al. (2025). Pancreatic cancer-restricted cryptic antigens are targets for T cell recognition.. Science. 388(6747):eadk3487. PMID: 40339010
- Weller C, Bartok O, et al. (2025). Translation dysregulation in cancer as a source for targetable antigens.. Cancer Cell. 43(5):823-840.e18. PMID: 40154482
- Huber F, Arnaud M, et al. (2025). A comprehensive proteogenomic pipeline for neoantigen discovery to advance personalized cancer immunotherapy.. Nature Biotechnology. 43(8):1360-1372. PMID: 39394480
- Cai Y, Gong M, et al. (2025). Immunopeptidomics-guided discovery and characterization of neoantigens for personalized cancer immunotherapy.. Science Advances. 11(21):eadv6445. PMID: 40397742
- Ishino T, Watanabe T, et al. (2026). Immunopeptidomics combined with full-length transcriptomics uncovers diverse neoantigens.. Cell Reports. 45(1):116781. PMID: 41447530
- Van der Vreken A, Thery F, et al. (2025). Immunopeptidomics identified antigens for mRNA-lipid nanoparticle vaccines with alpha-galactosylceramide in multiple myeloma therapy.. Journal for ImmunoTherapy of Cancer. 13(4):e010673. PMID: 40300855
- Yu KKH, Basu S, et al. (2025). Investigative needle core biopsies support multimodal deep-data generation in glioblastoma.. Nature Communications. 16(1):3957. PMID: 40295505
Recent Posts
-
The Neural Shield: How Cancer-Induced Nerve Injury Drives Immunotherapy Resistance
For years, our understanding of cancer focused primarily on the tumor cells themselves and the immu …29th Mar 2026 -
Metabolic Bloodhounds: Engineering Immune Cells to Track Cancer's Chemical Trail
Imagine a specialized squad of immune cells, meticulously trained and equipped, not just to fight c …27th Mar 2026 -
Mitochondrial Transplantation: A New Frontier in Restoring Neural Function After Ischemic Stroke
Imagine a future where the devastating impact of a stroke could be significantly mitigated, not jus …27th Mar 2026