Description
Human Prothrombin (F2) ELISA Kit
The Human Prothrombin (F2) ELISA Kit is specifically designed for the quantitative detection of prothrombin levels in human samples such as serum, plasma, and cell culture supernatants. This kit offers exceptional sensitivity and specificity, ensuring precise and consistent results for a variety of research purposes.Prothrombin, also known as coagulation factor II, is a key player in the blood clotting process, essential for stopping bleeding and maintaining hemostasis. Dysregulation of prothrombin levels is associated with various clotting disorders, making it a critical biomarker for studying clotting conditions and developing new treatments.
With the Human Prothrombin (F2) ELISA Kit, researchers can accurately measure prothrombin levels in human samples, opening up new possibilities for understanding coagulation disorders and potentially identifying novel therapeutic targets. Trust in the reliability and accuracy of this ELISA kit for your research needs.
| Product Name: | Human Prothrombin (F2) ELISA Kit |
| SKU: | HUEB0326 |
| Size: | 96T |
| Target: | Human Prothrombin (F2) |
| Synonyms: | Coagulation factor II |
| Assay Type: | Sandwich |
| Detection Method: | ELISA |
| Reactivity: | Human |
| Detection Range: | 0.78-50ng/mL |
| Sensitivity: | 0.32ng/mL |
| Intra CV: | 5.4% | ||||||||||||||||||||
| Inter CV: | 7.1% | ||||||||||||||||||||
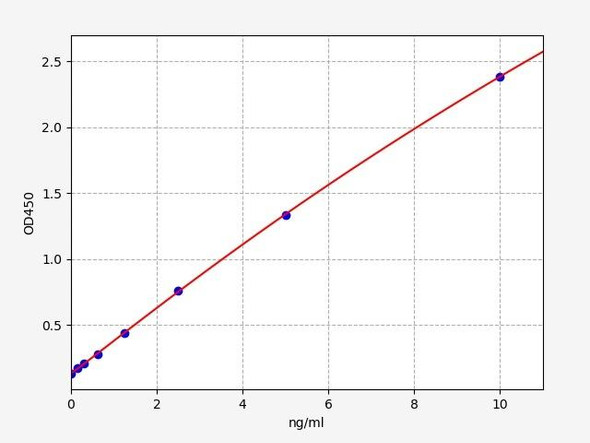
| Linearity: |
| ||||||||||||||||||||
| Recovery: |
| ||||||||||||||||||||
| Function: | Thrombin, which cleaves bonds after Arg and Lys, converts fibrinogen to fibrin and activates factors V, VII, VIII, XIII, and, in complex with thrombomodulin, protein C. Functions in blood homeostasis, inflammation and wound healing. |
| Uniprot: | P00734 |
| Sample Type: | Serum, plasma, tissue homogenates, cell culture supernates and other biological fluids |
| Specificity: | Natural and recombinant human Prothrombin |
| Sub Unit: | Heterodimer (named alpha-thrombin) of a light and a heavy chain; disulfide-linked. Forms a heterodimer with SERPINA5. |
| Research Area: | Cancer |
| Subcellular Location: | Secreted Extracellular space |
| Storage: | Please see kit components below for exact storage details |
| Note: | For research use only |
| UniProt Protein Function: | Function: Thrombin, which cleaves bonds after Arg and Lys, converts fibrinogen to fibrin and activates factors V, VII, VIII, XIII, and, in complex with thrombomodulin, protein C. Functions in blood homeostasis, inflammation and wound healing. Ref.12 |
| UniProt Protein Details: | Catalytic activity: Selective cleavage of Arg-|-Gly bonds in fibrinogen to form fibrin and release fibrinopeptides A and B. Subcellular location: Secreted extracellular space. Tissue specificity: Expressed by the liver and secreted in plasma. Post-translational modification: The gamma-carboxyglutamyl residues, which bind calcium ions, result from the carboxylation of glutamyl residues by a microsomal enzyme, the vitamin K-dependent carboxylase. The modified residues are necessary for the calcium-dependent interaction with a negatively charged phospholipid surface, which is essential for the conversion of prothrombin to thrombin. Involvement in Disease: Defects in F2 are the cause of factor II deficiency (FA2D) [ MIM:613679]. It is a very rare blood coagulation disorder characterized by mucocutaneous bleeding symptoms. The severity of the bleeding manifestations correlates with blood factor II levels. Ref.2 Ref.30 Ref.31 Ref.32 Ref.33 Ref.34 Ref.35 Ref.36 Ref.37 Ref.38 Ref.39 Ref.40Genetic variations in F2 may be a cause of susceptibility to ischemic stroke (ISCHSTR) [ MIM:601367]; also known as cerebrovascular accident or cerebral infarction. A stroke is an acute neurologic event leading to death of neural tissue of the brain and resulting in loss of motor, sensory and/or cognitive function. Ischemic strokes, resulting from vascular occlusion, is considered to be a highly complex disease consisting of a group of heterogeneous disorders with multiple genetic and environmental risk factors. Ref.43Defects in F2 are a cause of susceptibility to thrombosis (THR) [ MIM:188050]. It is a multifactorial disorder of hemostasis characterized by abnormal platelet aggregation in response to various agents and recurrent thrombi formation. Note=A common genetic variation in the 3-prime untranslated region of the prothrombin gene is associated with elevated plasma prothrombin levels and an increased risk of venous thrombosis. Ref.32 Ref.41 Pharmaceutical use: The peptide TP508 also known as Chrysalin (Orthologic) could be used to accelerate repair of both soft and hard tissues. Miscellaneous: Prothrombin is activated on the surface of a phospholipid membrane that binds the amino end of prothrombin and factors Va and Xa in Ca-dependent interactions; factor Xa removes the activation peptide and cleaves the remaining part into light and heavy chains. The activation process starts slowly because factor V itself has to be activated by the initial, small amounts of thrombin.It is not known whether 1 or 2 smaller activation peptides, with additional cleavage after Arg-314, are released in natural blood clotting.Thrombin can itself cleave the N-terminal fragment (fragment 1) of the prothrombin, prior to its activation by factor Xa.The cleavage after Arg-198, observed in vitro, does not occur in plasma. Sequence similarities: Belongs to the peptidase S1 family.Contains 1 Gla (gamma-carboxy-glutamate) domain.Contains 2 kringle domains.Contains 1 peptidase S1 domain. |
| NCBI Summary: | Coagulation factor II is proteolytically cleaved to form thrombin in the first step of the coagulation cascade which ultimately results in the stemming of blood loss. F2 also plays a role in maintaining vascular integrity during development and postnatal life. Mutations in F2 leads to various forms of thrombosis and dysprothrombinemia. [provided by RefSeq] |
| UniProt Code: | P00734 |
| NCBI GenInfo Identifier: | 135807 |
| NCBI Gene ID: | 2147 |
| NCBI Accession: | P00734.2 |
| UniProt Secondary Accession: | P00734,Q4QZ40, Q53H04, Q53H06, Q7Z7P3, Q9UCA1, B2R7F7 B4E1A7, |
| UniProt Related Accession: | P00734,Q15253,Q16505,Q69EZ7,Q69EZ8,Q86WA1,Q8TD58 |
| Molecular Weight: | 70,037 Da |
| NCBI Full Name: | Prothrombin |
| NCBI Synonym Full Names: | coagulation factor II (thrombin) |
| NCBI Official Symbol: | F2 |
| NCBI Official Synonym Symbols: | PT |
| NCBI Protein Information: | prothrombin; serine protease; OTTHUMP00000197117; OTTHUMP00000233661; prothrombin B-chain |
| UniProt Protein Name: | Prothrombin |
| UniProt Synonym Protein Names: | Coagulation factor II |
| Protein Family: | Prothrombin |
| UniProt Gene Name: | F2 |
| UniProt Entry Name: | THRB_HUMAN |
| Component | Quantity (96 Assays) | Storage |
| ELISA Microplate (Dismountable) | 8×12 strips | -20°C |
| Lyophilized Standard | 2 | -20°C |
| Sample Diluent | 20ml | -20°C |
| Assay Diluent A | 10mL | -20°C |
| Assay Diluent B | 10mL | -20°C |
| Detection Reagent A | 120µL | -20°C |
| Detection Reagent B | 120µL | -20°C |
| Wash Buffer | 30mL | 4°C |
| Substrate | 10mL | 4°C |
| Stop Solution | 10mL | 4°C |
| Plate Sealer | 5 | - |
Other materials and equipment required:
- Microplate reader with 450 nm wavelength filter
- Multichannel Pipette, Pipette, microcentrifuge tubes and disposable pipette tips
- Incubator
- Deionized or distilled water
- Absorbent paper
- Buffer resevoir
*Note: The below protocol is a sample protocol. Protocols are specific to each batch/lot. For the correct instructions please follow the protocol included in your kit.
Allow all reagents to reach room temperature (Please do not dissolve the reagents at 37°C directly). All the reagents should be mixed thoroughly by gently swirling before pipetting. Avoid foaming. Keep appropriate numbers of strips for 1 experiment and remove extra strips from microtiter plate. Removed strips should be resealed and stored at -20°C until the kits expiry date. Prepare all reagents, working standards and samples as directed in the previous sections. Please predict the concentration before assaying. If values for these are not within the range of the standard curve, users must determine the optimal sample dilutions for their experiments. We recommend running all samples in duplicate.
| Step | |
| 1. | Add Sample: Add 100µL of Standard, Blank, or Sample per well. The blank well is added with Sample diluent. Solutions are added to the bottom of micro ELISA plate well, avoid inside wall touching and foaming as possible. Mix it gently. Cover the plate with sealer we provided. Incubate for 120 minutes at 37°C. |
| 2. | Remove the liquid from each well, don't wash. Add 100µL of Detection Reagent A working solution to each well. Cover with the Plate sealer. Gently tap the plate to ensure thorough mixing. Incubate for 1 hour at 37°C. Note: if Detection Reagent A appears cloudy warm to room temperature until solution is uniform. |
| 3. | Aspirate each well and wash, repeating the process three times. Wash by filling each well with Wash Buffer (approximately 400µL) (a squirt bottle, multi-channel pipette,manifold dispenser or automated washer are needed). Complete removal of liquid at each step is essential. After the last wash, completely remove remaining Wash Buffer by aspirating or decanting. Invert the plate and pat it against thick clean absorbent paper. |
| 4. | Add 100µL of Detection Reagent B working solution to each well. Cover with the Plate sealer. Incubate for 60 minutes at 37°C. |
| 5. | Repeat the wash process for five times as conducted in step 3. |
| 6. | Add 90µL of Substrate Solution to each well. Cover with a new Plate sealer and incubate for 10-20 minutes at 37°C. Protect the plate from light. The reaction time can be shortened or extended according to the actual color change, but this should not exceed more than 30 minutes. When apparent gradient appears in standard wells, user should terminatethe reaction. |
| 7. | Add 50µL of Stop Solution to each well. If color change does not appear uniform, gently tap the plate to ensure thorough mixing. |
| 8. | Determine the optical density (OD value) of each well at once, using a micro-plate reader set to 450 nm. User should open the micro-plate reader in advance, preheat the instrument, and set the testing parameters. |
| 9. | After experiment, store all reagents according to the specified storage temperature respectively until their expiry. |
When carrying out an ELISA assay it is important to prepare your samples in order to achieve the best possible results. Below we have a list of procedures for the preparation of samples for different sample types.
| Sample Type | Protocol |
| Serum | If using serum separator tubes, allow samples to clot for 30 minutes at room temperature. Centrifuge for 10 minutes at 1,000x g. Collect the serum fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. If serum separator tubes are not being used, allow samples to clot overnight at 2-8°C. Centrifuge for 10 minutes at 1,000x g. Remove serum and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. |
| Plasma | Collect plasma using EDTA or heparin as an anticoagulant. Centrifuge samples at 4°C for 15 mins at 1000 × g within 30 mins of collection. Collect the plasma fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. Note: Over haemolysed samples are not suitable for use with this kit. |
| Urine & Cerebrospinal Fluid | Collect the urine (mid-stream) in a sterile container, centrifuge for 20 mins at 2000-3000 rpm. Remove supernatant and assay immediately. If any precipitation is detected, repeat the centrifugation step. A similar protocol can be used for cerebrospinal fluid. |
| Cell culture supernatant | Collect the cell culture media by pipette, followed by centrifugation at 4°C for 20 mins at 1500 rpm. Collect the clear supernatant and assay immediately. |
| Cell lysates | Solubilize cells in lysis buffer and allow to sit on ice for 30 minutes. Centrifuge tubes at 14,000 x g for 5 minutes to remove insoluble material. Aliquot the supernatant into a new tube and discard the remaining whole cell extract. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20 °C. |
| Tissue homogenates | The preparation of tissue homogenates will vary depending upon tissue type. Rinse tissue with 1X PBS to remove excess blood & homogenize in 20ml of 1X PBS (including protease inhibitors) and store overnight at ≤ -20°C. Two freeze-thaw cycles are required to break the cell membranes. To further disrupt the cell membranes you can sonicate the samples. Centrifuge homogenates for 5 mins at 5000xg. Remove the supernatant and assay immediately or aliquot and store at -20°C or -80°C. |
| Tissue lysates | Rinse tissue with PBS, cut into 1-2 mm pieces, and homogenize with a tissue homogenizer in PBS. Add an equal volume of RIPA buffer containing protease inhibitors and lyse tissues at room temperature for 30 minutes with gentle agitation. Centrifuge to remove debris. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20 °C. |
| Breast Milk | Collect milk samples and centrifuge at 10,000 x g for 60 min at 4°C. Aliquot the supernatant and assay. For long term use, store samples at -80°C. Minimize freeze/thaw cycles. |